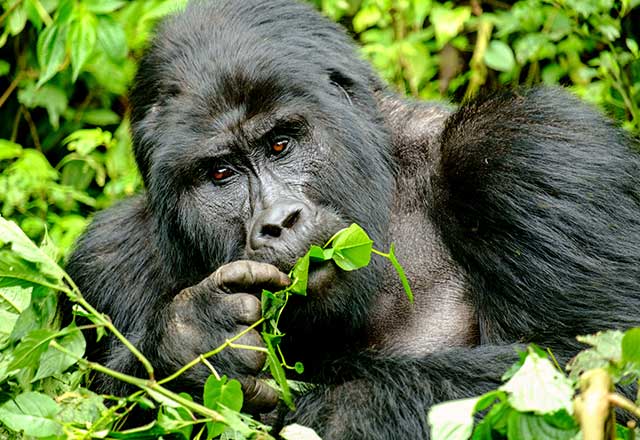
From the Trees to the Ground: What Gorilla Feet Can Teach Us About Evolution
Johns Hopkins researchers found that the heel bone looks different in gorillas who walk on land compared with those who live in the trees, establishing a new avenue in evolutionary and behavioral research.